Upload your small RNAs Next-Generation Sequencing data to detect and profile sncRNAs:
- Inputs could be collapsed FASTA files (Recommend) (with Fasta head as >A_xB,where A is unique seq name while B is the frequency of A), FASTQ format, GSM/SRR sample IDs or accessible links to FASTA files
- To speed the uploading, we highly recommend that the upload file is further compressed in .zip or .gz format
- If you large number of samples, use this Link to upload multiple samples in collapsed FASTA format.
FASTQ format
The raw data file should be in FASTQ format (See the figure below as example)
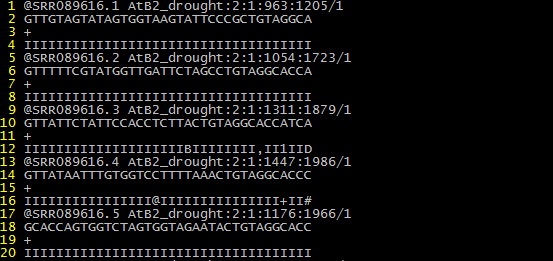
To further minimize the size of uploading file, users are highly recommended to use gzip (for Linux and Mac OS) to compress the fasta file into gz format file or use winrar (for Windows) to compress the fasta file into zip format file.
