Welcome to RiboMicrobe
RNA translation directly regulates protein synthesis and the translation process of microorganisms is essential for their vital functions, affecting their adaptability and survival in different environments. With the advancement of ribosome profiling (Ribo-seq), researchers are paying increasing attention to different RNA translation processes as well as microproteins encoded by small ORF (sORF) in various prokaryotic microorganisms.
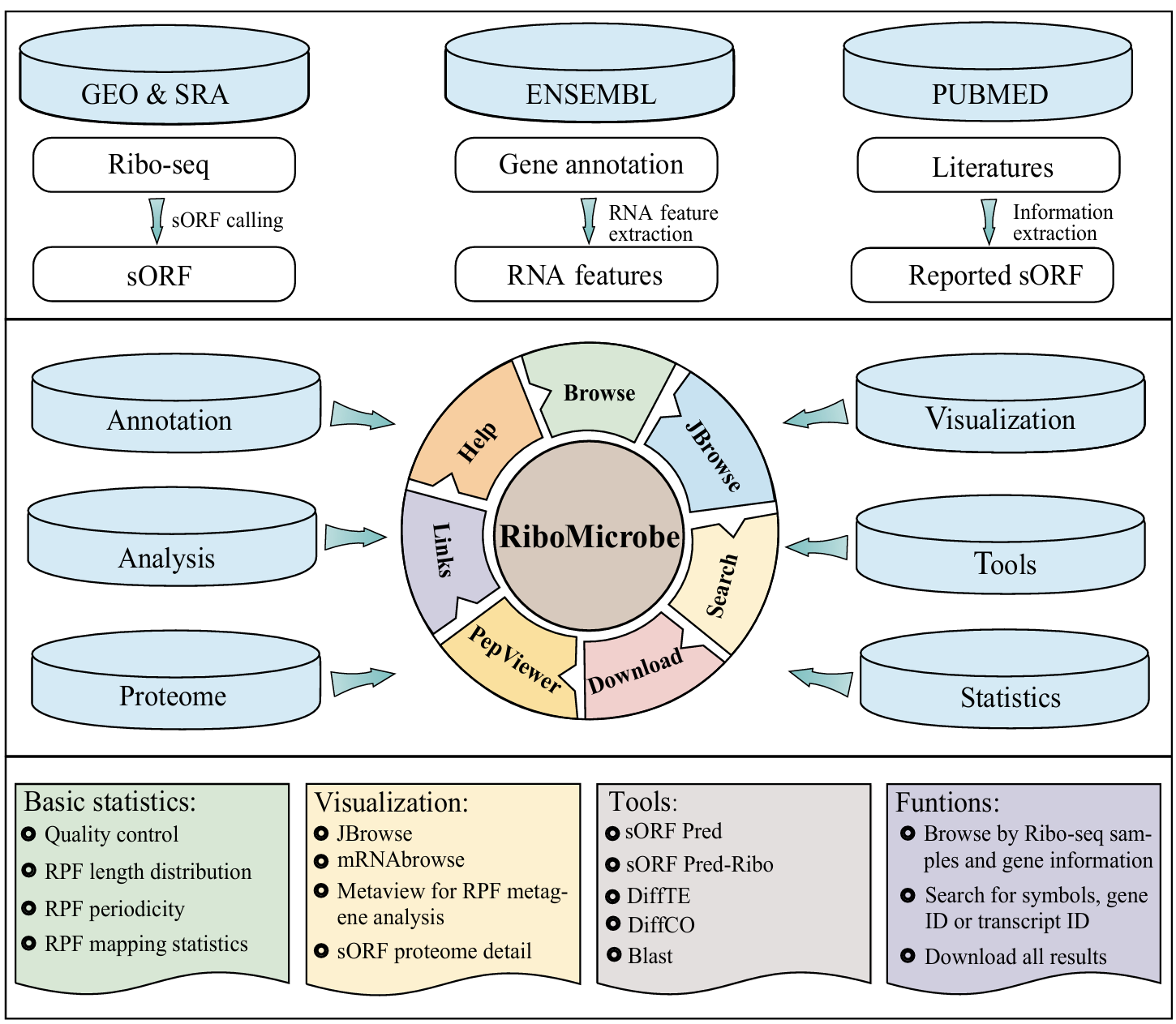
There is already very large number of translatome studies and public Ribo-seq data available for microorganisms. However, there is still no database of translatomics available for prokaryotic microorganism yet. Therefore, we present the RiboMicrobe database, which focus on extensive and integrate translation analyses of prokaryotic microorganism based on Ribo-seq data. To our best of knowledge, RiboMicrobe is the first comprehensive database which includes 38 different prokaryotic microorganism species, 891 Ribo-seq datasets, 369 associated RNA-seq datasets, 75 proteome datasets to extensively show different translatome characteristics and sORF-encoded micropeptides among different microorganism. Besides, we collected additional related datasets from previous research articles and related resources, such as diversify types of RNA modifications (m1A, m5C, m6A, m7G and pseudouridine) and riboswitches that can affect RNA translation in microorganism. Additionally, RiboMicrobe provided various useful functionalities and visualization tools, such as mRNAbrowse showing the transcript profile of potential sORF, JBrowse visualizing all ribo-seq samples and other publicly available annotations in genome scale, PepViewer showing the mass spectrum information of sORF, diffTE for differentially translation analysis, diffCodonOccupancy for different codon occupancy analysis and blast for sORF homology analysis.
The output of RiboMicrobe includes:
- The overall statistic of RPF features across different species
- The overall statistic of different gene features and biotypes
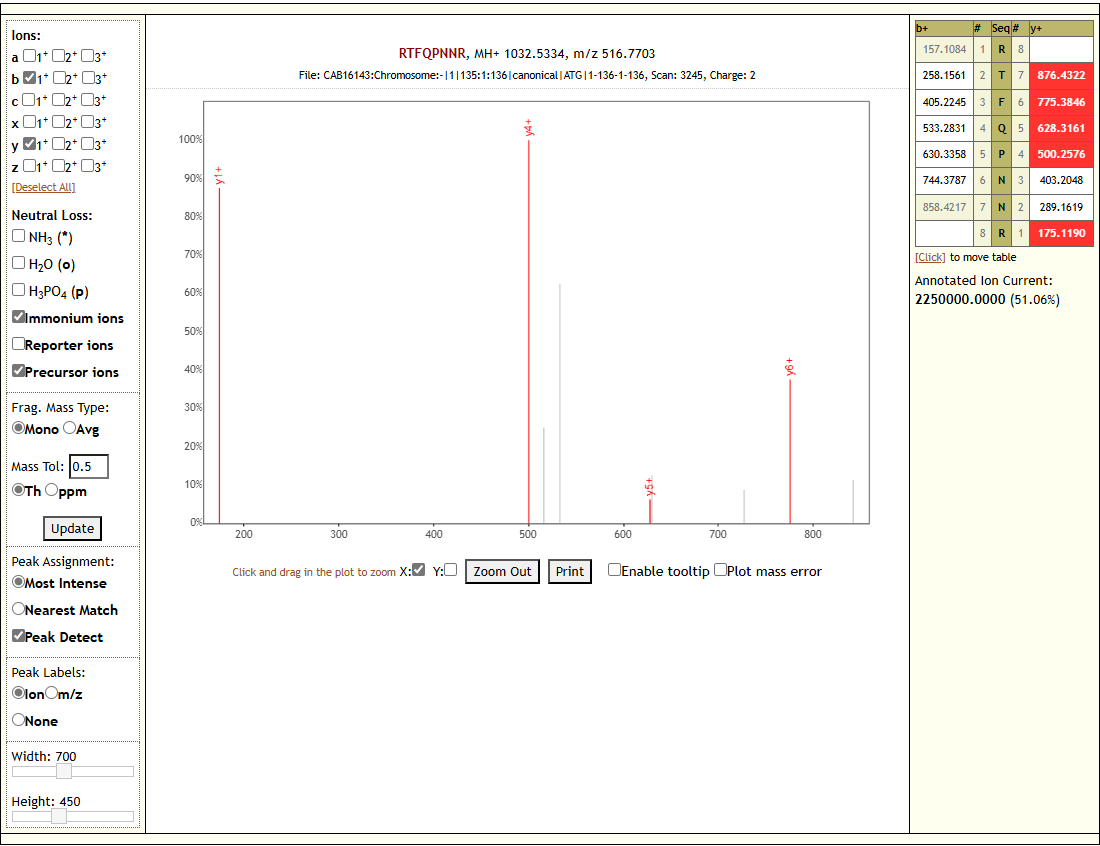
- The Proteomics analysis visualizes the identified peptides
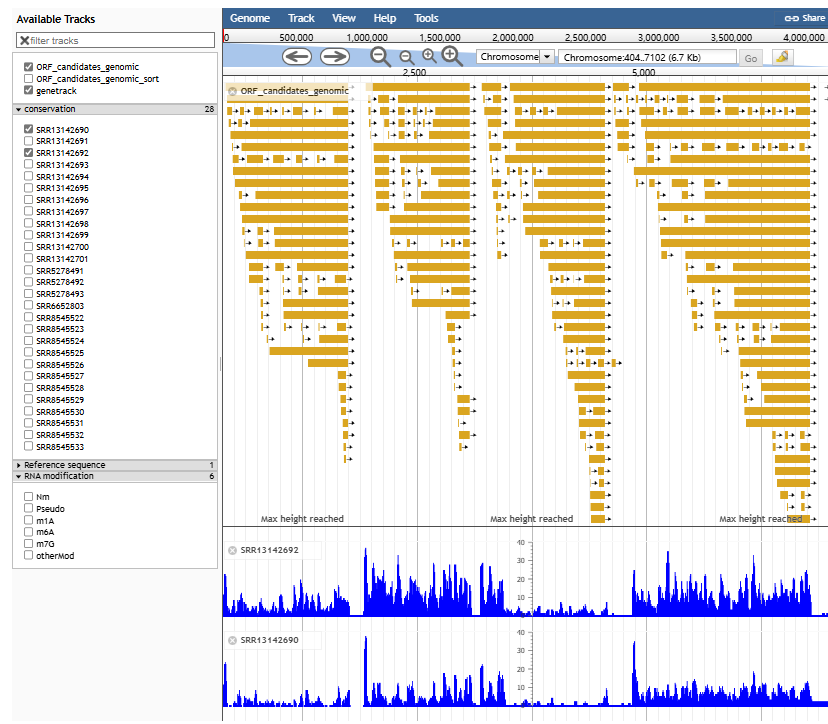
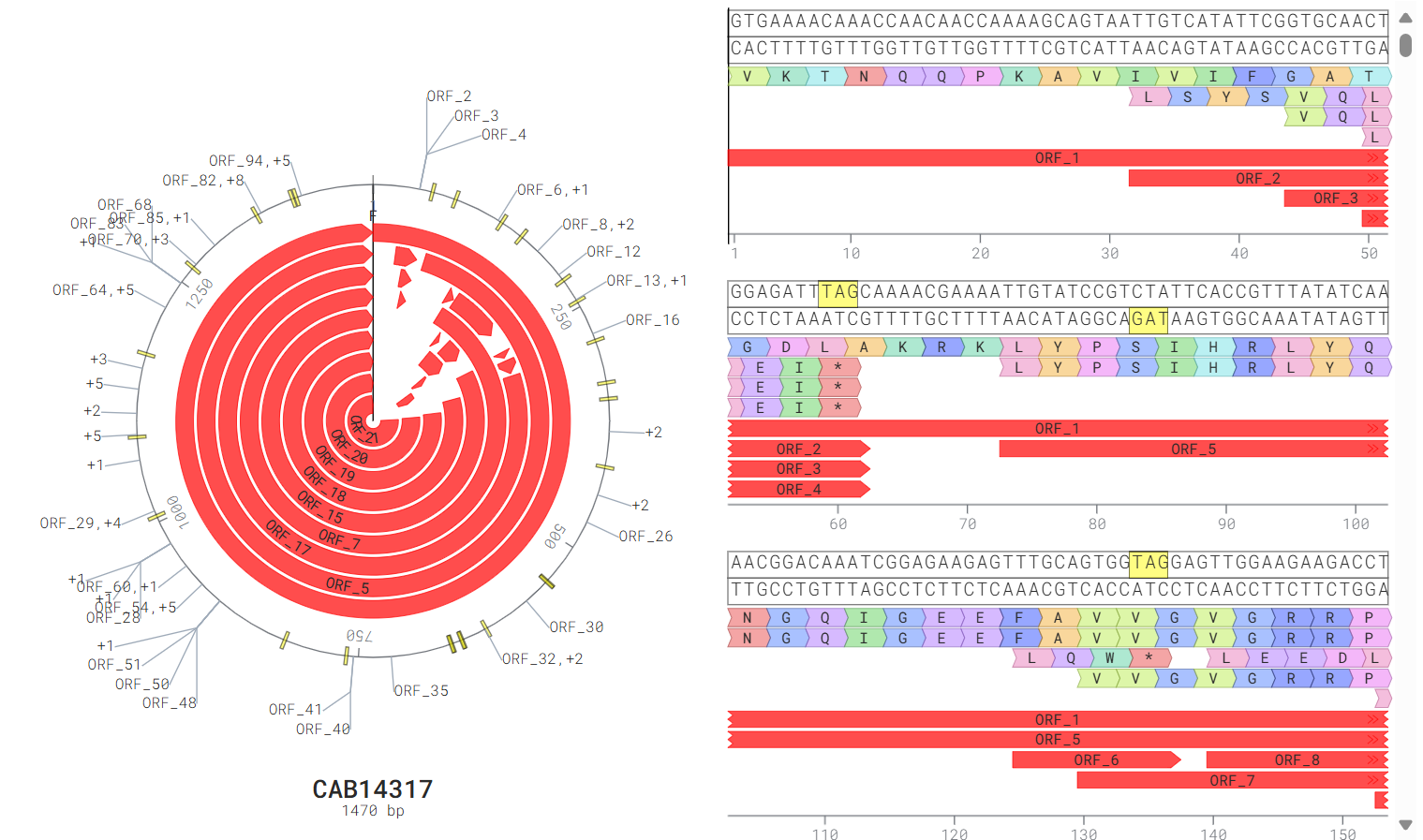
- JBrowse and mRNAbrowse are integrated to visualize the gene feature
- sORFPredRibo and sORFPred are integrated to predict potential sORFs.
- The DiffTE tool to identify differences in translation efficiency between samples
- The DiffCO tool to identify differences in codon occupancy between samples
- The Blast tool for search short peptide
RiboMicrobe: an integrated translatome atlas for microorganism
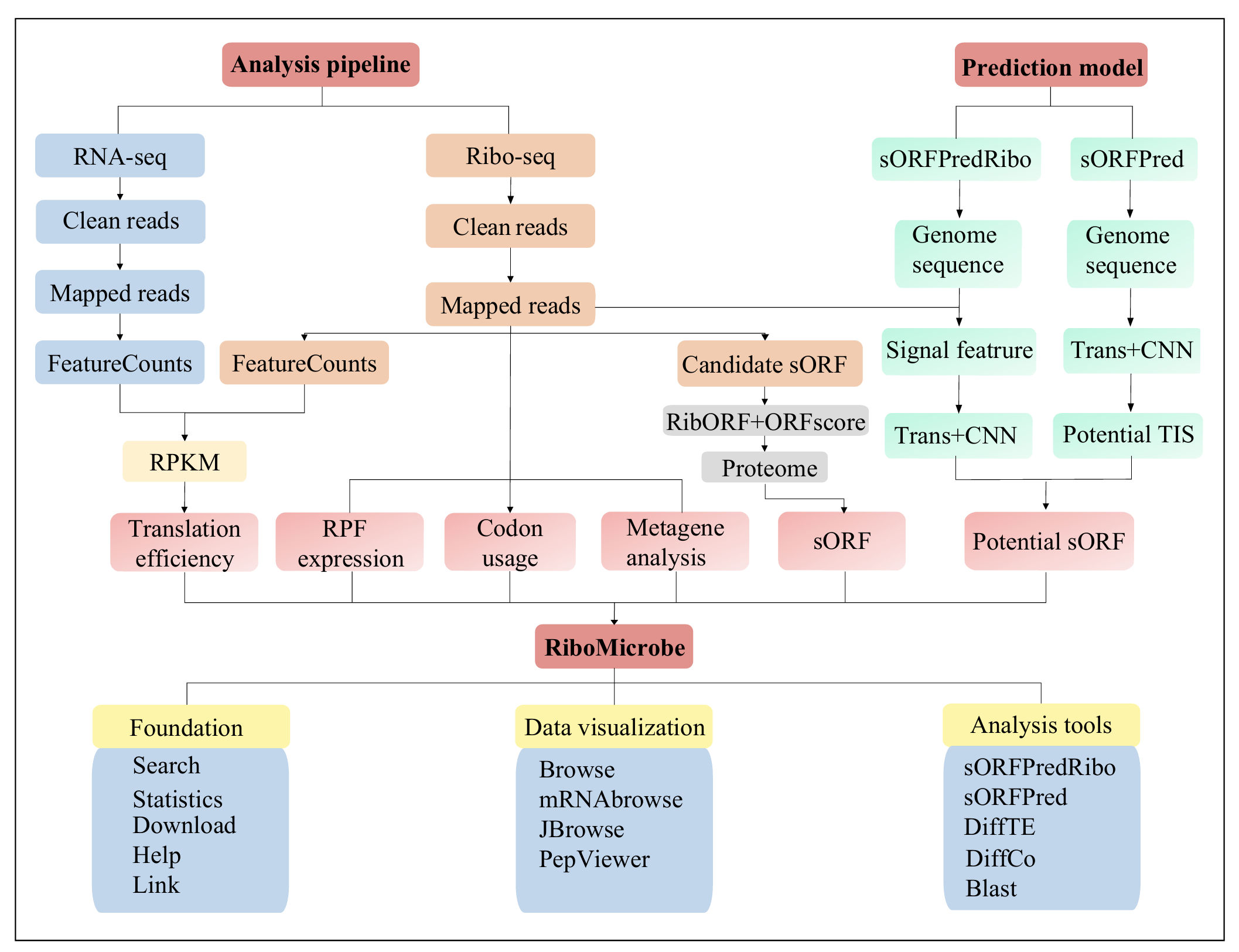
Data Visualization
Analysis Tools
Recent news More
Help is upioading.
Home is upioading.
PepViewer is upioading.
sORFPredRibo is developed in RiboMicrobe.